cuBNM
A toolbox for efficient simulation of brain biophysical network models on GPUs
Cite this software
Description
cuBNM toolbox uses GPUs to efficiently run simulations of brain network models consisting of nodes which are connected through a connectome, and fit them to empirical neuroimaging data through integrated optimization algorithms.
GPU parallelization enables massive scaling of the simulations into higher number of simulations and nodes. Below you can see how computing time varies as a function of number of simulations and nodes on GPUs versus CPUs. For example, running 32,768 simulations (duration: 60s, nodes: 100) would take 3.8 days on a single CPU thread, but only 5.6 minutes on Nvidia A100 GPU, and 21.8 minutes on Nvidia GeForce RTX 4080 Super:
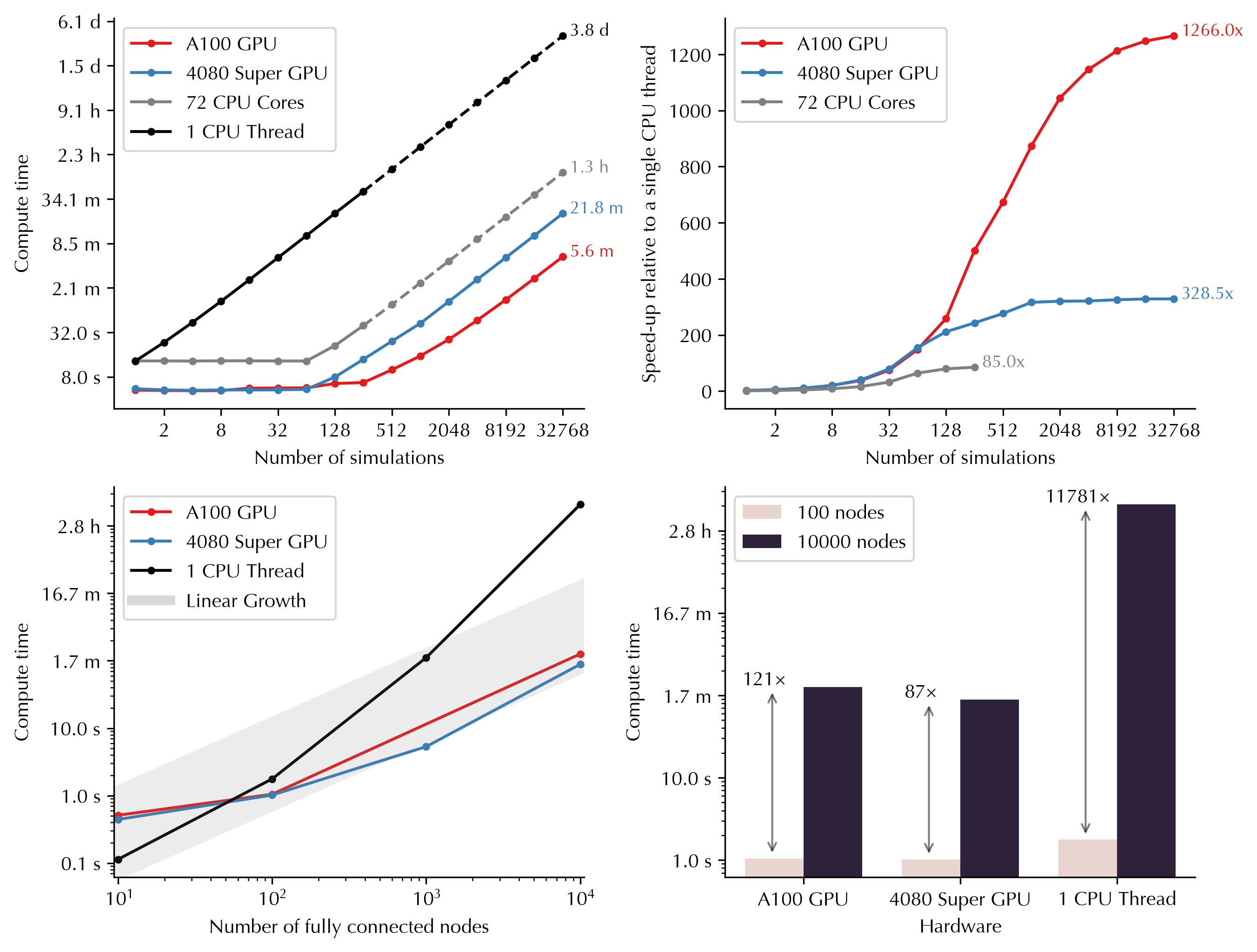
GPU usage is the primary focus of the toolbox but it also supports running the simulations on single or multiple cores of CPU. CPUs will be used if no GPUs are detected or if requested by the user.
Several commonly used models (e.g., reduced Wong-Wang, Jansen-Rit, Kuramoto, Wilson-Cowan) are implemented, and new models can be added via YAML definition files. A guide is included on the structure of model definition YAML files to help researchers implement custom models.
The simulated activity of model neurons is fed into the Balloon-Windkessel model to calculate simulated BOLD signal. Functional connectivity (FC) and functional connectivity dynamics (FCD) from the simulated BOLD signal are calculated efficiently on GPUs/CPUs and compared to FC and FCD matrices derived from empirical BOLD signals to assess similarity (goodness-of-fit) of the simulated to empirical BOLD signal. The toolbox supports parameter optimization algorithms including grid search and evolutionary optimizers (via pymoo), such as the covariance matrix adaptation-evolution strategy (CMA-ES). Parallelization within the grid or the iterations of evolutionary optimization is done at the level of simulations (across the GPU ‘blocks’), and nodes (across each block’s ‘threads’). The models can incorporate global or regional free parameters that are fit to empirical data using the provided optimization algorithms. Regional parameters can be homogeneous or vary across nodes based on a parameterized combination of fixed maps or independent free parameters for each node or group of nodes.
At its core the toolbox runs highly parallelized simulations using C++/CUDA, while the user interface is written in Python and allows for user control over simulation configurations:
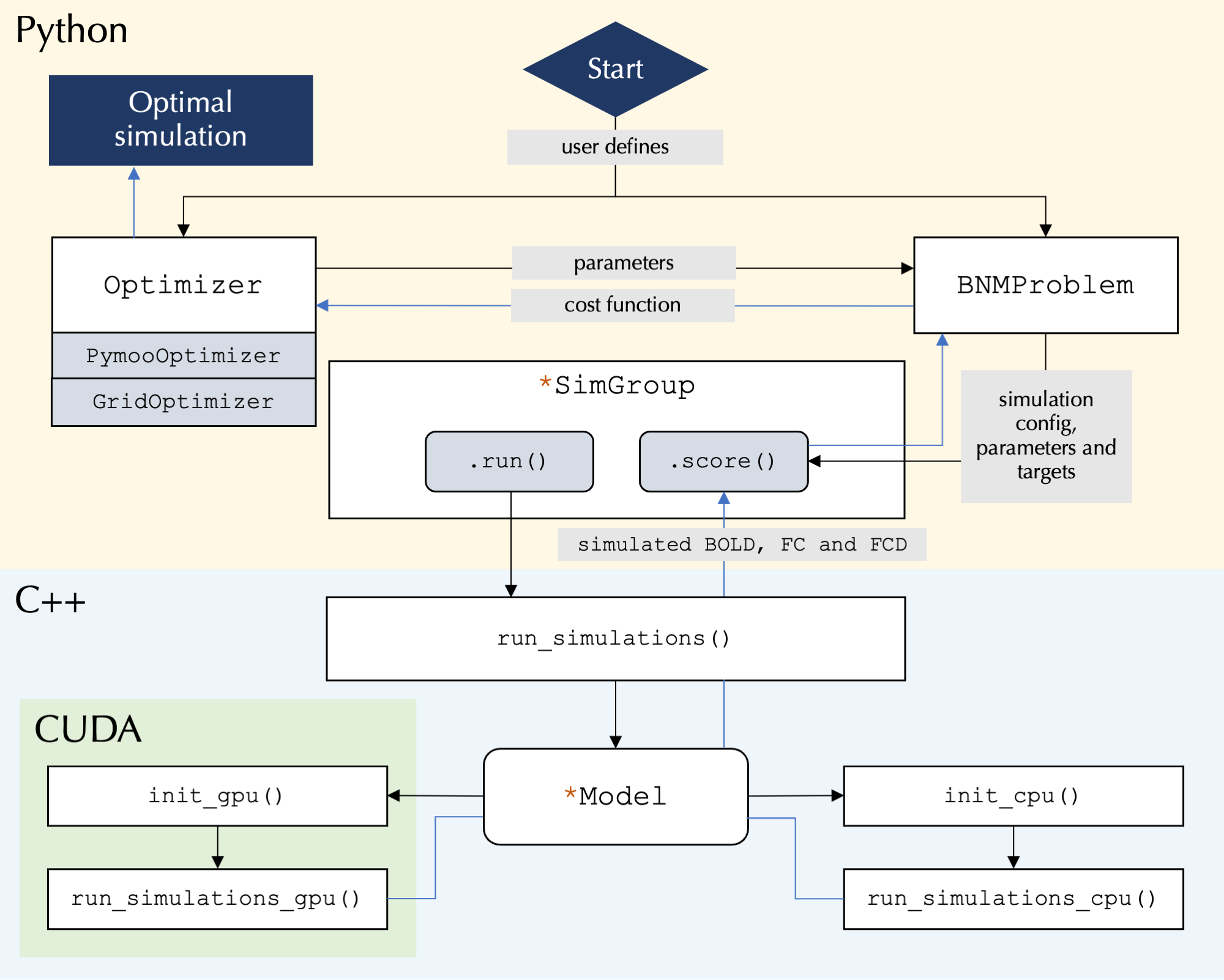

Participating organisations
Reference papers
Mentions
- 1.Author(s): Jessica Royer, Casey Paquola, Sara Larivière, Justine Y. Hansen, Sofie L. Valk, Bratislav Misic, Robert Leech, Jonathan Smallwood, Boris C. BernhardtPublished in Nature Reviews Neuroscience by Springer Science and Business Media LLC in 202610.1038/s41583-026-01038-0
- 2.Author(s): Flávia A Verza, Ícaro S Freitas, Enzo P Valenzuela, Anthony A Grace, Francisco S Guimarães, Felipe V GomesPublished in Cerebral Cortex by Oxford University Press (OUP) in 202610.1093/cercor/bhaf342
- 3.Author(s): Francesca Saviola, Stefano Tambalo, Laura Beghini, Asia Ferrari, Barbara Cassone, Dimitri Van De Ville, Jorge JovicichPublished in Proceedings of the National Academy of Sciences by Proceedings of the National Academy of Sciences in 202510.1073/pnas.2506513122
- 4.Author(s): Jason Z. Kim, Richard F. Betzel, Ahmad Beyh, Amber Howell, Amy Kuceyeski, Bart Larsen, Caio Seguin, Xi-Han Zhang, Avram Holmes, Linden ParkesPublished in Nature Communications by Springer Science and Business Media LLC in 202510.1038/s41467-025-66542-w
- 5.Author(s): Valerie J. Sydnor, Amar Ojha, Bart Larsen, Angela Martinez, Finnegan J. Calabro, Beatriz LunaPublished in Neuropsychopharmacology by Springer Science and Business Media LLC in 2025, page: 67-8510.1038/s41386-025-02246-5
- 6.Author(s): Yuehua Xu, Tengda Zhao, Mingrui Xia, Xuhong LiaoPublished in Journal of Pacific Rim Psychology by SAGE Publications in 202510.1177/18344909251392757
- 7.Author(s): Ruizhe Liu, Hyesang Chang, Yuan Zhang, Vinod MenonPublished in Developmental Cognitive Neuroscience by Elsevier BV in 2025, page: 10162810.1016/j.dcn.2025.101628
- 8.Author(s): Guoshi Li, Khoi Minh Huynh, Kim-Han Thung, Hoyt Patrick Taylor IV, Guoye Lin, Weili Lin, Sahar Ahmad, Pew-Thian YapPublished in Communications Biology by Springer Science and Business Media LLC in 202510.1038/s42003-025-08970-4
- 1.Author(s): Alejandro Orozco Valero, Natalia Kopiś-Posiej, Víctor Rodríguez-González, Víctor Gutiérrez-de Pablo, Christian Morillas, Jesús Poza, Carlos Gómez, Pablo Martínez-Cañada, Paweł KrukowPublished by openRxiv in 202610.64898/2026.01.14.699432
- 2.Author(s): Victor Barnes, Jace Cruddas, Trang Cao, Isaac Z. Pope, Ting Xu, Thomas Funck, Nicola Palomero-Gallagher, James C. Pang, Alex FornitoPublished by openRxiv in 202610.64898/2026.01.22.701178
- 3.Author(s): Ghazal Mirzaee, Roshanak Latifikhereshki, Jonathan Chang, Shahrzad LatifiPublished in 202610.1016/j.mlwa.2026.100905